-
On March 9, 2022, researchers from the Institut Pasteur used the global data science initiative GISAID (1-3) to release a severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) genome (ID: EPI_ISL_10819657) from an isolated virus and announced that it was the first solid evidence for a recombinant strain from 2 types of variants of concern (VOCs) of SARS-CoV-2 (lineage AY.4 and BA.1, belonging to Delta and Omicron, respectively). Complementary to the original data submitters’ full analysis, we would like to comment on this case in context of our experience in coronavirus evolution and the further perspective of this finding on the course of the pandemic. As announced, this novel strain has high genomic similarity to viruses belonging to lineage AY.4, except for the region encoding the spike (S) gene, which is more similar to those from lineage BA.1 (Figure 1A). Therefore, this novel strain uses Delta as its genomic backbone and then replaces a large portion of original S gene with the ortholog from Omicron. A total of 36 amino acid changes were found in the S protein compared to the prototype of SARS-CoV-2 (Figure 1B). Among 36 amino acid mutations, 27 were found in BA.1, 5 mutations were found in AY.4, and 4 mutations were found in both AY.4 and BA.1. However, it was not the first case of a recombination event identified in SARS-CoV-2. A previous study has documented that inter-lineage recombination events have been found in SARS-CoV-2 and then some recombinants caused further community transmission (4). However, these inter-lineage recombination events only occurred in some loci of the genome. No recombination events involving large genomic fragments (like “Deltacron”) have been found in SARS-CoV-2 before.
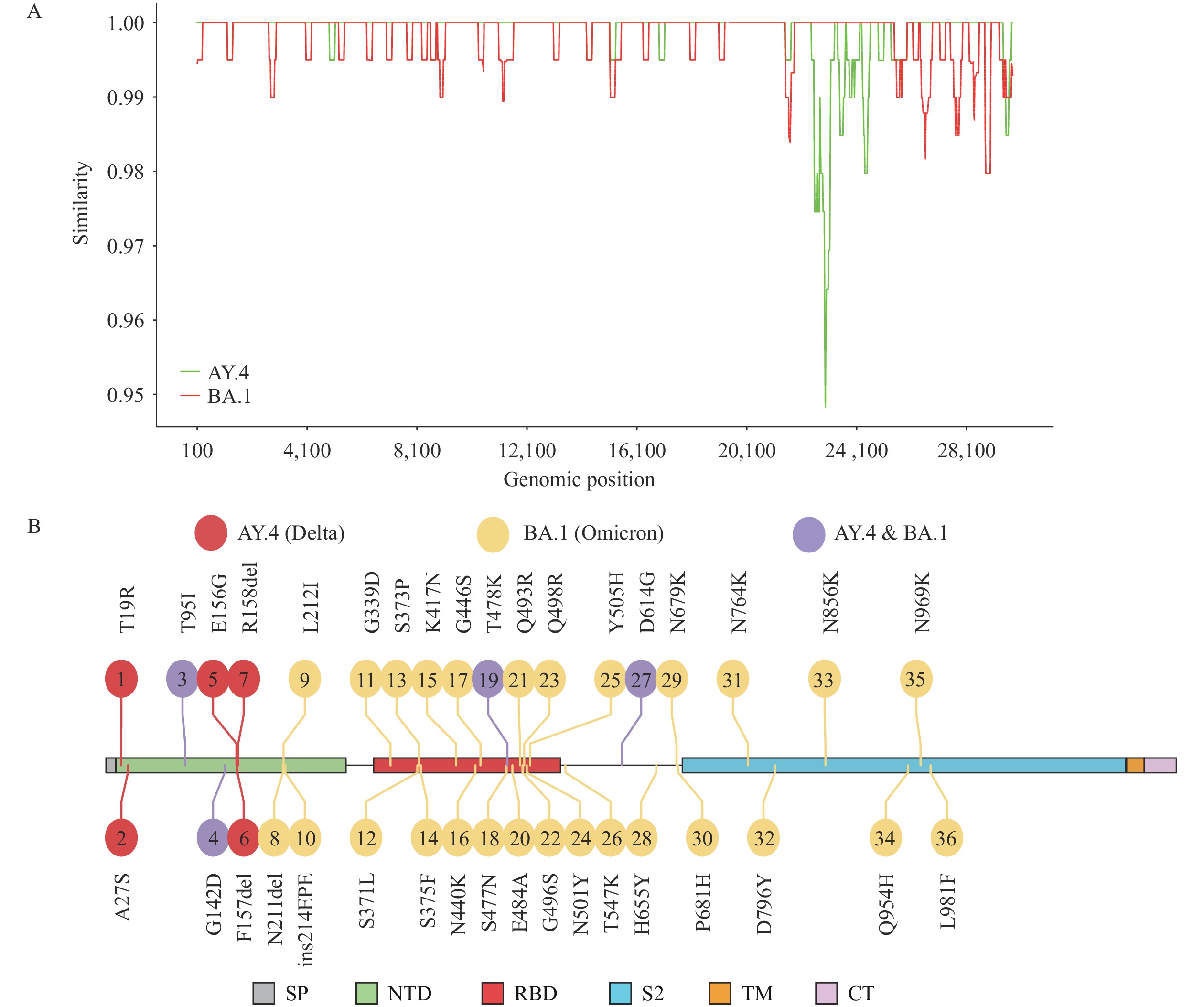 Figure 1.
Figure 1.Genetic variation of “Deltacron.” (A) The nucleotide similarity of “Deltacron” compared to AY.4 and BA.1; (B) The mutation profile of the S protein encoded by “Deltacron.”
Note: Mutations in only AY.4, BA.1, both AY.4 and BA.1, and “Deltacron” were assigned to different colors. Abbreviations: SP=signal peptide; NTD=N terminal domain; RBD=receptor binding domain; TM=transmembrane domain; CT=C terminal. -
Coronaviruses (CoVs) belong to a highly diverse family Coronaviridae, which could infect numerous types of species (like those from Aves and mammalian). Therefore, CoVs will pose a potential risk to public health and economy. As a hallmark of CoVs, genetic recombination events occur frequently in natural reservoir hosts due to co-infection of different types of CoVs in a single individual host and/or other reasons. Genetic recombination could arise at both the intra-species and inter-species level. For intra-species recombination, some genetic materials could be exchanged between strains of different subtypes of the same species. For example, an isolate of the Middle East respiratory syndrome coronavirus (MERS-CoV), imported to China from the Republic of Korea, has been documented as a recombinant virus from group 3 and group 5 (5). On the other hand, inter-species recombination occurs when two different species exchange their partial genetic materials and are also common in nature (including several types of human CoVs) (6). Despite most recombination events occurring among species belonging to Coronaviridae, cross-family genetic recombination events had also been found in nature (between Coronaviridae and Reoviridae) (7). The frequent genetic recombination of CoVs could lead to the emergence of novel viruses. Further, the most important threat of the emergence of these novel recombinant viruses is the possibility of cross-species transmission (8). The most recent example was two canine-feline recombinant alphacoronaviruses with extremely high genomic similarity (99.4%) were isolated from humans by two independent research groups in different countries (9-10).
-
Before the emergence of “Deltacron,” most scientific efforts were focused on assessing and responding to the effects of point mutations in the genome of SARS-CoV-2 during its global spread and evolution. Until now, 5 types of SARS-CoV-2 VOCs had been found circulating globally (Alpha, Beta, Gamma, Delta, and Omicron). They all exhibited increased transmissibility and varying degrees of immune escape (11-12). Although the emergence and subsequent spread of these VOCs has had a huge impact on global health and economy, it may not have been the worst case until now, as recombination (a major mechanism bringing genetic diversity to coronaviruses) had not really emerged on a large scale and shown its power before the emergence of “Deltacron.” The emergence of “Deltacron” is therefore a “grey rhino” event rather than a “black swan” event.
-
With the advent of “Deltacron,” further concerns are coming. According to our preliminary analysis on the first “Deltacron” genome, recombination event only occurs in S gene. However, recombination events involving all types of genes encoded by CoVs have been found in nature (13). Consequently, the genetic recombination events involving other genes and/or the combination of other genes would also occur with high probability. Furthermore, the genetic recombination event of “Deltacron” occurs at the inter-lineage level, that is the parents of the “Deltacron” came from different lineages of the same species (AY.4 and BA.1), respectively. We could speculate that cross-species recombination events would also appear in future for the following reasons. First, the variety of host types of CoVs has resulted in its wide distribution throughout the world (14) and SARS-CoV-2 has spread all over the world. Second, several species of mammals other than humans have been infected by SARS-CoV-2 in nature, and many more species have been shown to be susceptible to SARS-CoV-2 (15). In addition, the spillover events of SARS-CoV-2 between humans and animals in both directions (for example transmission of SARS-CoV-2 from humans to minks and then back to humans and further community transmission) (16-17) have been found in nature. Taken together, the probability of co-infection with SARS-CoV-2 and other types of CoVs or even other viruses in a single host would be high, leading to the occurrence of cross-species recombination with high probability. Therefore, it is difficult to predict which viral species SARS-CoV-2 will recombine with, and on which genes future recombination will occur. This kind of uncertainty is doomed to increase the likelihood of generating a novel recombinant virus with unknown risk to humans.
-
Although the emergence of a recombinant SARS-CoV-2 isolate was expected by scientists, it still attracted great attention. In addition to the need to conduct in-depth evaluation and research on the various properties of this novel recombinant virus, and adjust in prevention and control strategies based on the results, it is more important to be alert to the generation of other types of recombinant viruses produced by SARS-CoV-2 and other viruses. Therefore, it is particularly important to implement large-scale virome study in both domesticated and wild animals.
-
The data contributors, i.e., the authors from the originating laboratory (Labo Analyses Med) and the Submitting laboratory (Etienne Simon-Loriere and colleagues in National Reference Center for Viruses of Respiratory Infections, Institut Pasteur, Paris), GISAID, and Peter Bogner.
HTML
| Citation: |




 Download:
Download:




