-
Dengue fever (DF) is a major threat to human life and affects a large number of populations in the tropical and subtropical zones with serious consequences. Over 300 million people are infected worldwide with dengue virus (DENV), which causes DF, every year per the World Health Organization (WHO) (1). Since 1978, DF outbreaks occurred endemically and reached epidemic levels every 4 to 7 years in China, and increased significantly in recent decades in Guangdong Province (2). According to China Information System for Disese Control and Prevention, DF outbreaks occurred from May to November in Guangdong in 2019, and the reported number of cases started to increase rapidly in August, peaking in October, and decreasing by November (Figure 1). As of November 30, a total of 5,968 cases, including 3 deaths, were reported to the Guangdong Provincial CDC. Among them, 1,207 cases were cases imported from abroad, and 4,761 cases were acquired through presumed local mosquito-borne transmission. The cities with the most reported cases were Guangzhou (1,376), Shantou (914), Foshan (375), Zhanjiang (336), Chaozhou (288), Jiangmen (256), Jieyang (246), Zhongshan (239), Shenzhen (208), Dongguan (116), and Qingyuan (110); all other cities in Guangdong Province had less than 100 reported cases.
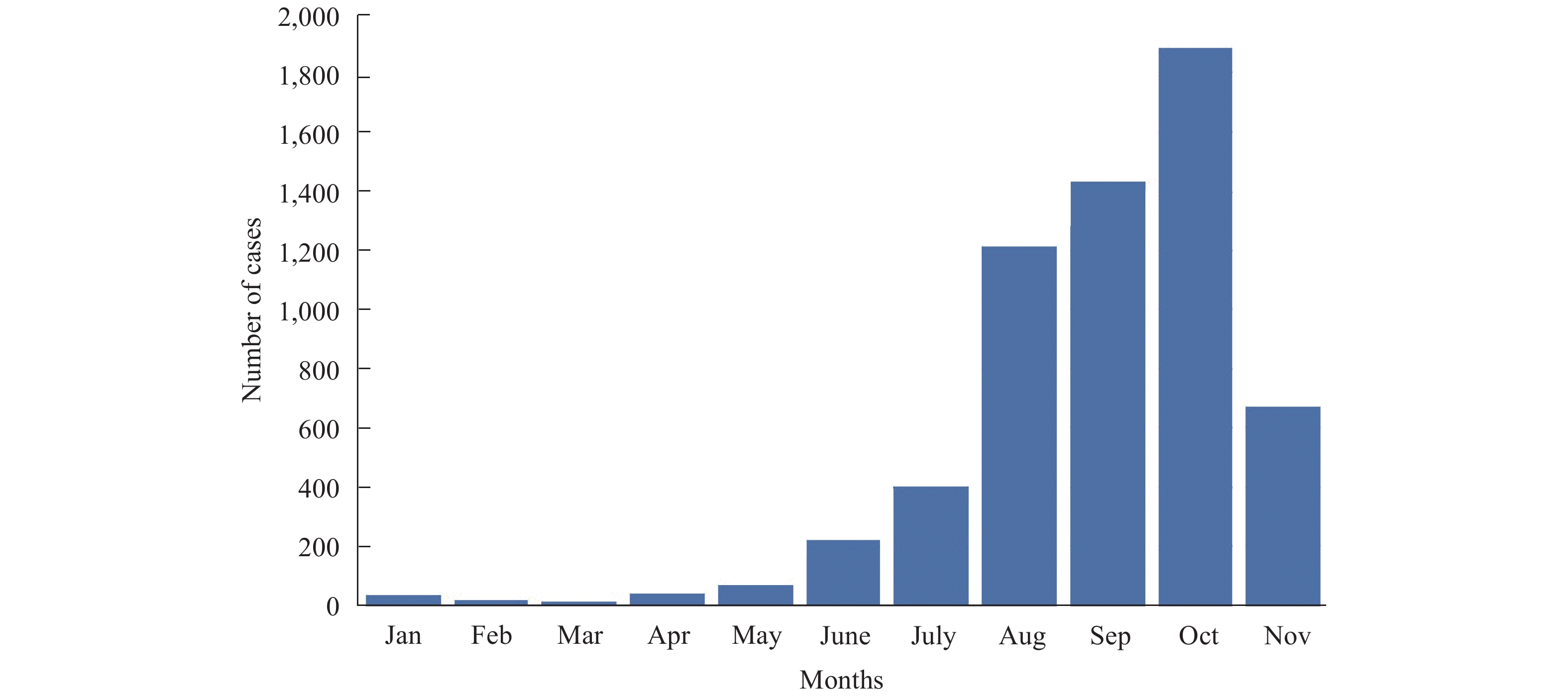 Figure 1.
Figure 1.Number of dengue fever cases reported from January to November in 2019 in Guangdong Province, China.
In this study, 21 city-level CDCs were involved in collection of serum specimens of suspected DF cases from respective local hospitals, and these samples were sent to Guangdong Provincial CDC for laboratory detection and further analysis. As of November 20, a total of 2,293 serum specimens were collected, detected, and serotyped using real-time RT-PCR. A total of 2,175 samples were positive for dengue virus, and the serotyping results showed that 1,919 samples were type I, 159 were type II, 93 were type III, and 4 were type IV DENV infection. Virus isolation was performed using RT-PCR to detect positive samples using cell culture technology, and 427 strains were successfully isolated including 359 strains of type I, 47 strains of type II, 18 strains of type III, and 3 strains of type IV.
To trace the origins of DENV circulating in Guangdong, the entire E gene of DENV samples were sequenced from imported cases, sporadic cases acquired through presumed local transmission, and representative virus strains obtained in outbreaks from different period in 2019. A total of 326 sequences were successfully obtained, and the US National Library of Medicine Basic Local Alignment Search Tool (BLAST) from the US National Center for Biotechnology Information (NCBI) analysis showed that 287 sequences belong to type I DENV (75 from imported cases and 212 from local acquired cases), 27 type II (16 imported and 11 local), 10 type III (7 imported and 3 local), and 2 type IV sequences from imported cases.
Phylogenetic analysis of the obtained E gene sequences were performed with 5 genotype representative sequences, which demonstrated that the 287 type I E gene sequences were clustered into 2 different genotypes, 280 sequences clustered to genotype I and 7 clustered to genotype V. The 212 sequences of genotype I DENV E gene, obtained from cases acquired through presumed local transmission, were clustered into 6 distinct clades, and clade 6 contained 190 sequences followed by clade 4 containing 9 sequences, clade 3 containing 8 sequences, clade 5 containing 3 sequences, and clade 1 and clade 2 containing 1 sequences. The sequences from clade 6 and clade 3 were clustered together with viral sequence obtained from cases acquired in Cambodia and shared the highest homology at the nucleotide level, which suggested that sustained local transmission might be caused by virus introduced from Cambodia. Clade 1 and clade 2 formed an independent branch, and they might be caused by virus introduced from Thailand and Myanmar. Clade 4 shared the highest homology at nucleotide level with strains imported from Thailand (D19014), and clade 5 shared the highest homology at nucleotide level with strains imported from Malaysia (D16167). In addition, the sequences of genotype V were clustered together with strains imported from Indonesia (D19011). These results suggested that DF epidemics in Southeast Asia put great pressure on DF control and prevention in Guangdong and that Cambodia, Thailand, East Timor, Malaysia, and Indonesia were sources of DENV introduced into Guangdong in 2019. The dominant genotype of the virus in subsequent local transmission was genotype I of type I DENV, which circulated in more than 16 cities, and genotype V of type I DENV was mainly found in outbreaks occurred in Zhongshan and Guangzhou (Figure 2).
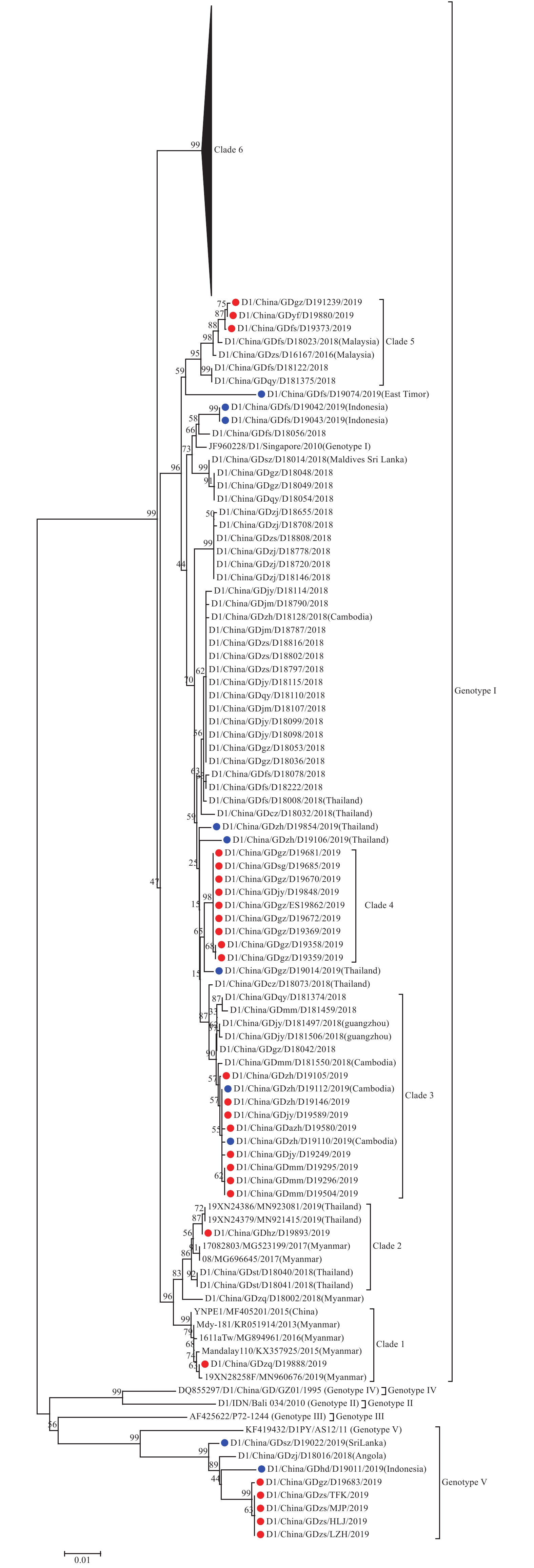 Figure 2.
Figure 2.Phylogenetic tree of type-1 dengue virus isolated from the 2019 outbreak in Guangdong Province, China. A total of 287 E gene sequences from isolated strains were used for phylogenetic tree reconstructions, and 5 genotypes were assigned according to previous studies. The outbreak isolates are indicated by red circles, and the isolates from the imported cases are indicated by blue circles.
Type II and III viruses were found among cases acquired both abroad and through local transmission, while DENV of type IV was only detected in imported cases. A total of 27 sequences of type II DENV E gene were clustered into Asia I and cosmopolitan genotypes, 10 sequences of cosmopolitan genotypes from cases acquired through local transmission were clustered together with viruses circulating in Cambodia and in Malaysia in clade 1 and clade 2, which revealed a possible source of the viruses causing local transmission in Guangdong. The sequence of genotype Asia I of local transmitted type II DENV clustered together with viruses introduced from Vietnam. The 10 sequences of DENV type III viruses clustered into genotype I and II respectively, among which two local strains belonged to genotype I, which clustered into the same clade with the virus imported from Indonesia, and the sequence of local transmitted virus of genotype II showed high similarity with imported strains from Cambodia.
-
In the history of Guangdong (including Hainan Province), more than 10,000 cases of DF have been reported in six years, and the latest one was 2014, with more than 45,000 cases were reported (3-4). In recent years, the frequency of DF outbreaks increased in Guangdong and the affected area has expanded year after year. Before 2011, dengue fever was prevalent in Guangdong for only one to two months, and then gradually increased to seven to eight months. Outbreaks of DF have also started occurring yearly since 2011, ranging from thousands to tens of thousands of cases. The affected area had expanded from being concentrated in the Pearl River Delta to widely affecting most of Guangdong Province. This may be related to the gradual rise of the earth’s temperature and the rapid spread of DENV, but it could also be related to the rapid development of China’s economy, the acceleration of urbanization, and increasing foreign trade and tourism (5-6).
The annual circulation of DENV has expanded from a single type of DENV to multiple serotypes and genotypes circulating simultaneously. DENV type I has caused outbreaks every year since 2011; type II has caused outbreaks every year since 2013, accounting for the highest proportion in 2015 and reaching 63.82% of all cases; type III has caused local outbreaks in 2013, 2015, 2016, and 2019 (7-8); type IV virus was mainly detected from imported cases with only a few local transmission caused by type IV virus were reported. In 2019, the dominant type of dengue virus in Guangdong was type I, and type II and III were also detected in outbreaks through local transmission.
DF outbreaks in Guangdong were still often local outbreaks caused by imported viruses. Cambodia was the main source of DENV introduced to Guangdong in 2019, followed by Indonesia, Thailand, and Malaysia. According to data from China Information System for Disese Control and Prevention, imported dengue viruses from more than 43 countries and regions were found in Guangdong in 2019, and Southeast Asia was the most important source, of which Cambodia contributed the largest number of imported cases, followed by Thailand and Malaysia. The surveillance results are consistent with our molecular tracing results in this report, suggesting that DF outbreaks in Guangdong were mainly caused by imported virus. Therefore, early detection of suspected cases and typing and genotyping of circulating viruses should be prioritized and enhanced to promptly assess the likelihood of local transmission, of introduction, and of subsequent sustained local transmission of the virus to implement optimal prevention and control strategies and measures.
Acknowledgments: We acknowledge all of the staff at the 21 CDCs for their contributions to sample collection and surveillance data collection in Guangdong.
HTML
| Citation: |




 Download:
Download:




