-
Human norovirus (NoV) is the leading cause of epidemics of viral acute gastroenteritis (AGE) worldwide, affecting people in all age groups. Data showed that NoVs were genetically diverse and played an increasingly important role in the etiology of AGE in China (1), among which GII.2[P16] recombinant NoV reemerged and caused outbreaks in some Asian countries like China and Japan in 2016. The mechanism behind the sudden epidemic is poorly characterized. In this study, we summarized and analyzed the major potential reasons for the re-emergence and the prevalence of GII.2[P16] NoV.
-
NoVs are non-enveloped, single-stranded, positive sense, polyadenylated RNA viruses that belong to the Norovirus genus, Caliciviridae family. The genome is approximately 7.3–7.7 kb in length and consists of three open reading frames (ORFs): ORF1 encodes a polyprotein that is further cleaved into six nonstructural proteins (P48, NTPase, P22, VPg, Pro, and RdRp); ORF2 encodes the major capsid protein VP1 and can be divided into shell (S) and protruding (P) domains, and the P domain is further subdivided into P1 and P2 subdomains, where P2 is a hypervariable region, determining the antigenicity and cell binding of the virus; ORF3 encodes the minor structural protein VP2. The genomes begin with a 5' end terminal pGpU sequence that is covalently linked to VPg, and a short-conserved region (CR) at the 5' end is repeated internally in the genome near the beginning of a subgenomic-sized RNA transcript (Figure 1).
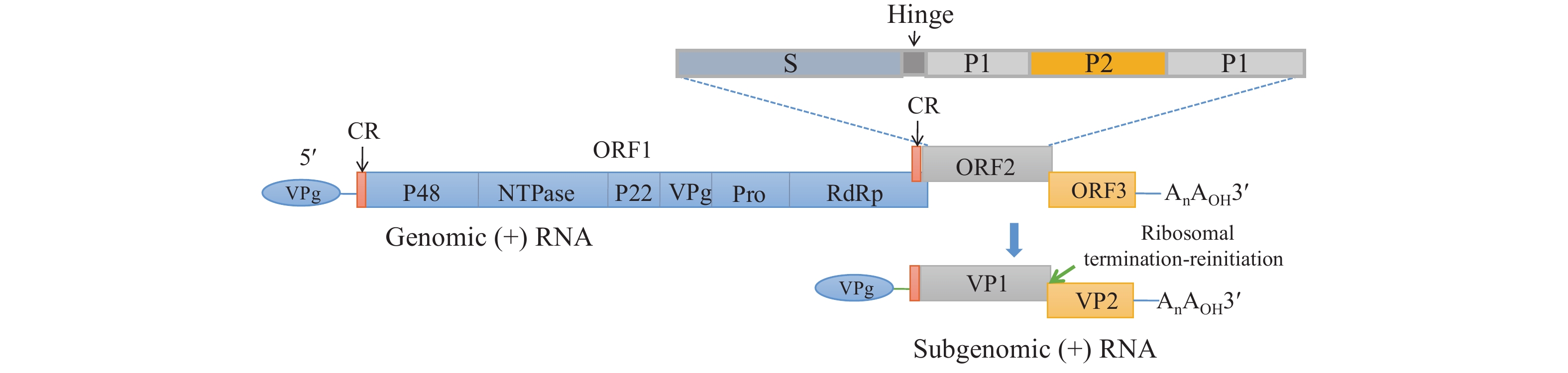 Figure 1.
Figure 1.NoV genome structure and the encoding proteins.
Note: ORF1 encodes a polyprotein further cleaved into six nonstructural proteins. ORF2 encodes the major capsid protein and can be divided into S and P domains — the P domain is further subdivided into P1 and P2 subdomains. ORF3 encodes the minor structural protein. Abbreviations: CR=conserved region; ORF=open reading frame; RdRp=RNA dependent RNA polymerase; S=shell domain; P=protruding domain.NoVs have high genetic and antigenic diversity. Based on amino acid diversity of the complete VP1 and nucleotide diversity of the RdRp, NoVs can be divided into 10 (GI–GX) genogroups with 48 confirmed genotypes and 60 confirmed P-types (2). GII genogroup viruses are the most commonly detected in humans; they can be further divided into 27 confirmed genotypes and 37 P types. From the mid-1990s to 2014, GII.4 genotype and its new variants have caused about 70%−80% of all NoV-associated AGE outbreaks worldwide (3). However, non-GII.4 NoVs have also severely affected China and some other Asian countries; in 2014 to 2015, GII.17 NoV emerged and increased in prevalence, while since 2016, GII.2[P16] reemerged and became predominant in AGE outbreaks.
-
Though the GII.2 genotype has been reported since the 1970s, it was only detected in sporadic cases and accounted for less than 2% of all NoV genotypes. GII.2[P16] recombinant NoV first appeared in Japan in 2008 and caused AGE outbreaks in Osaka at that time and then was occasionally detected in sporadic cases. In 2016, GII.2[P16] NoV reemerged in Guangdong Province, China, and rapidly became the main epidemic strain in Asian countries (4-6). The occurrence of GII.2[P16] from sporadic to large-scale outbreaks suggested that the reemergence and sudden epidemic may be related to the change of viral biological properties, which led to stronger transmissive and infective ability.
Multiple amino acid changes at antigenic sites have always been the most important factors for the emergence of an antigenically distinct GII.4 virus (7). However, compared with the GII.2 viruses from before 2016, the currently prevalent GII.2[P16] has no unique amino acid changes on VP1, and phylogenetic analyses revealed that capsid protein did not play a role in the potential of GII.2 NoV to become an epidemic (8). It was also shown that different GII.2 strains circulating between 1976 and 2010 had undergone a limited amount of evolution in blockade epitopes (9). All these results revealed that the reemergence of GII.2[P16] NoV was not caused by the antigenic drift of capsid proteins. On the contrary, compared with GII.P16 RdRp detected before 2016, five unique amino acid substitutions (D173E, S293T, V332I, K357Q, and T360A) were found in the novel GII.P16 RdRp; except for V332I, all were located on the surface of the protein. Since a single amino acid change on the RdRp surface of GII.4 NoV can affect the biological function (10), it is speculated that the unique amino acid changes on RdRp of GII.2[P16] could have certain impacts on the viral replication activity. Indeed, it was observed that the viral load of current prevalent GII.2[P16] NoV was higher than those of GII.4 and GII.17 genotypes in the whole population (11), suggesting that the enhanced replication ability was an important factor for the recurrence of GII.2[P16] NoV.
RdRp is a key protein that determines the replication efficiency of the NoV genome, and amino acid substitutions on it may alter the replication kinetics or fidelity of the virus. However, besides GII.2, the emerging novel GII.P16 RdRp has been recombined with seven other different capsid genotypes: GII.1, GII.3, GII.4, GII.12, GII.13, GII.16, and GII.17. A total of 312 nearly full genome sequences of these different genotypes were downloaded from NCBI GenBank database (http://www.ncbi.nlm.nih.gov/genbank) in this study. Amino acid comparison results showed that, except for GII.13, GII.16 and GII.17 still recombining with GII.P16 RdRp before 2016, the capsids from the 5 other genotypes after 2016 were recombined with the novel GII.P16 RdRp (Table 1). However, among these 5 different novel recombinant genotypes, GII.2[P16] caused 81.2% of the NoV-related AGE outbreaks in China (12), indicating that the novel GII.P16 RdRp was not the only factor that enhanced the viral replication ability. As a matter of fact, previous studies have determined that NoV proteins P48, VP1, and VP2 can modulate GII.4 RdRp activity in a species-specific manner (13). Compared with the GII.2[P16] before 2016, excluding RdRp, the current GII.2[P16] had 9 unique amino acid sustitutions in other nonstructural proteins. Amino acids in position 78-79 in P48, 147 in P22, and 49 in Pro of the viruses that had the novel GII.P16 RdRp were all EE, Q, and I. Except GII.12, the amino acids in position 165 of P48, 312 of NTPase, and 52 of P22 on the viruses that had novel GII.P16 RdRp were all R, P, and R, respectively (Table 1). This phenomenon suggested that the P48, NTPase, P22, and Pro together with the novel GII.P16 RdRp may enhance the viral replication ability and the fitness to the host.
As described above, VP1 can enhance the RdRp activity of NoV, we inferred that GII.P16 RdRp recombined with different types of capsid may potentially have different effects on the replication ability of the recombinant viruses, which needs further detailed scientific support. Furthermore, compared with the rare prevalent genotypes, the long-term prevalent ones like GII.4 would have a better fitness and stronger transmission ability in the population, and indeed, GII.4 [P16] was the primary cause of NoV outbreaks in the United States, Canada, and many other non-Asian countries at present (14-15). The regional specificity of GII.2[P16] and GII.4[P16] has little relation to the biological properties of the virus itself, while it may be attributed to the differences in the population’s genetic background and lifestyes, the climate, the environment, and so on.
-
A previous study suggested that the non-GII.4 genotype NoVs were static with their capsid as they were unable to evolve antigenically, and they would be prevalent for only a short period in limited areas and populations and then would shift to another genotype (8). Until now, GII.2[P16] NoV has been the predominantly circulating strain in China and in many Asian countries for more than five years since its reemergence in 2016. The evolutionary patterns for the viruses are unclear. Will the virus disappear after a period of epidemic? Or will the virus evolve into new variants that can lead to new outbreaks by adapting to new susceptible populations and drive escape from herd immunity every 2–7 years just like GII.4 did? Or will the virus generate new receptor-binding sites like GII.13 and GII.21 did? Or will the virus optimize receptor-binding sites for enhancing receptor-binding ability just like GII.17 did? Or even, will the virus acquire another novel RdRp and produce novel recombinant just like it already did? So, it is important to trace the genomic signatures and dynamic evolution of the GII.2[P16] NoV, which can provide scientific evidence and guidance for the prevention and control of the virus.
Genotypes/Variants P48 NTPase P22 Pro RdRp 52 78-79 165 312 52 147 158 49 173 293 332 357 360 GII.2 2009−2014 N − K A K P V V D S V K T GII.2 2010−2012 N E- K S K P A V D S V K T GII.2 2016−2019* K† EE§ R† P† R† Q§ T† I§ E§ T§ I§ Q§ A§ GII.4 2013 NA NA K P R S V V D S V K T GII.4 2016−2019* E/G† EE§ R† P/S† R† Q§ A/T† V/I E§ T§ V/I Q§ A§ GII.3 2011−2015 N E- K P/S K P A V D S V K T GII.3 2016−2018* E† EE§ R† P/S† R† Q§ T† I§ E§ T§ I§ Q§ A§ GII.1 2016−2018* K/E† EE§ K/R† P† R† Q§ P/S¶ I§ E§ T§ I§ Q§ A§ GII.12 2017−2018* K† EE§ K¶ S† K¶ Q§ P¶ I§ E§ T§ I§ Q§ A§ GII.13 2011−2015 N E- K S K P A/T V D S V K T GII.13 2016−2018* N/K E-/EE K/R P/S K/R P/Q A/T V/I D/E S/T V/I K/Q T GII.16 2018* N − K A K P A V D S V K T GII.16 2012 N E- K S K S A V D S V K T GII.17 2014 N E- K A K P T V D S V K T Note: “−” stands for amino acid deletion; “/” stands for “or,” e.g., R/K stands for R or K in the site.
Abbreviations: RdRp=RNA dependent RNA polymerase; NoV=norovirus; NA=not applicable.
* The GII.P16 NoVs currently prevalent.
† Key amino acid sites with changes on the currently circulating NoVs that had the novel GII.P16 RdRp.
§ Unique amino acids in currently predominant viruses.
¶ Different amino acids between currently circulating GII.[P16] strains.Table 1. Amino acid comparisons of nonstructural proteins between different NoV genotypes/variants recombined with GII.[P16] RdRp.
HTML
Genomic Feature and Genetic Diversity of NoV
Potential Factor for the Reemergence of GII.2[P16] in China
Potential Evolutionary Pattern of GII.2[P16] NoV in the Future
| Citation: |



 Download:
Download:




