-
After no local cases had been reported for 110 consecutive days in Tianjin Municipality, a confirmed case of coronavirus disease 2019 (COVID-19) was reported on June 17, 2020 in a 22-year-old male with no notable medical conditions. A total of 136 cases had previously been confirmed within Tianjin. The patient was employed as a kitchen assistant in the western dining room of a 5-star hotel, and within a 14-day period before onset of illness, he had not traveled to any other provincial-level administrative divisions (PLADs) or countries and had no known contact with suspected or confirmed cases, overseas returnees, or persons with similar respiratory symptoms cases. The patient developed fever (maximum temperature 38 °C), dry throat, and slight cough beginning on the morning of June 16, 2020 at approximately 8:00 am. He was transferred to Haihe Hospital for isolation and treatment after a COVID-19 nucleic acid test was positive on June 17. He was diagnosed as a COVID-19 confirmed case and was classified as mild by an expert group.
-
To prevent further spread of the disease and find the source of infection, Tianjin Municipal CDC organized relevant professionals to conduct epidemiological investigations and laboratory tests to identify the source of infection as rapidly as possible. Persons included in this investigation were the 137th confirmed COVID-19 patient in Tianjin, his family members and colleagues, residents of his apartment building, and hotel guests including non-Chinese nationals at his place of employment.
The patient was investigated with the “Epidemiological Questionnaire for COVID-19 Cases,” and others were investigated with a checklist that included basic personal information, clinical conditions, travel history, contact with others, and suspected exposures in the previous 14 days (1-3). Type and frequency of contact with the patient were determined.
Throat swabs and blood samples were obtained from all investigated subjects. Environmental samples from the patient’s residence and workplace were obtained including samples from door handles, faucets, kitchenware, refrigerators, garbage cans, air conditioners, sewers, clothes, stoves, sealed meat products, and aquatic equipment in the hotel. Samples from the environment, food, and food packaging from hotel suppliers were also obtained for testing.
Nucleic acid samples were extracted from throat, nasopharyngeal, and anal swabs using viral DNA/RNA extraction kits (Xi’an Tianlong Science and Technology, China), and initially stored at –70 °C. COVID-19 virus (also known as SARS-CoV-2 and 2019-nCoV) detection was conducted via real-time reverse transcriptase polymerase chain reaction (RT-PCR) using nucleic acid detection kits (Shanghai BioGerm Medical Technology, China) according to the manufacturer instructions. ULSEN® SARS-CoV-2 whole genome capture kits (MicroFuture, Beijing, China) were used for COVID-19 virus genome enrichment as described by the manufacturer. Amplified products were purified by Agencourt AMPure XPKit (Beckman Coulter, Brea, CA, USA) and then subjected to library preparation by Nextera® XT DNA Library Preparation Kit (Illumina, San Diego, CA, USA). Whole genome sequencing was carried out using a MiniSeq™ Sequencing System (Illumina, San Diego, CA, USA).
Resulting sequences were assembled with CLC Genomics Workbench 20.0.3 (Qiagen, Hilden, Germany). Sequence alignments and phylogenetic analysis were performed with MEGA X software, and a neighbor-joining tree was assembled with bootstrap values determined from 1,000 replicates. All comparison sequences, including the most recent Beijing outbreak sequences (i.e., ESP_ISL_469254, ESP_ISL_469255, and ESP_ISL_469256), were downloaded from the EpiFlu database of the Global Initiative on Sharing All Influenza Data (GISAID; gisaid.org) and from GenBank (ncbi.nlm.nih.gov/genbank/).
A total of 246 environmental and food specimens were collected during the investigation. This total included 144 environmental samples of 18 types of sources (e.g., door handles, kitchenware items, etc.) and 102 food samples (i.e., food and food packaging for frozen and fresh seafood, beef, mutton, and vegetables). All 246 specimens had negative viral nucleic acid test results.
A total of 1,137 contacts were investigated for suspected exposure (e.g., family members, colleagues, neighbors, hotel guests, etc.). No individuals had travel history to Xinfadi Agricultural Product Wholesale Market in Beijing Municipality or other high-risk areas or contact with persons with travel history to mid- to high-risk areas in Beijing. No individual had travel history overseas or contact with foreign travel returnees. In addition, no individual had contact with confirmed COVID-19 cases or persons experiencing COVID-19 or related respiratory symptoms.
On June 17, a total of 750 throat swab specimens and 365 serum specimens from persons with suspected exposures were collected and tested for both viral nucleic acids and virus-specific antibodies. All test results were negative.
On June 19, 55 employees in the dining room of the hotel were tested a second time. All viral nucleic acid test results were negative. All antibody test results were also negative, except one. A chef who worked in the western dining room of the hotel tested positive for IgM antibody specific to COVID-19 virus. After three consecutive tests, the IgM antibody result remained positive.
As a result, the chef who tested positive for anti-COVID-19-virus IgM antibodies was then further investigated as a possible source, and he reported having had no symptoms and was therefore preliminarily designated as a confirmed asymptomatic COVID-19 patient. He had visited his girlfriend in Beijing many times in the past month. While in Beijing on June 8, he took the subway and spent time in public places including restaurants, bars, and a theater. While in the workplace in the hotel in Tianjin, the chef and the patient had several exposures while in the smoking room on the ground floor. In addition, there was also an instance of close (less than 0.5 meters) communication between the chef and the patient on June 10 where the patient wore a mask but the chef did not. The patient was also responsible for cleaning the tableware in the employee’s dining room and had resulting contact with the chef during the daily lunch. Therefore, it was hypothesized that the source may have been infected in public places in Beijing, returned to work normally due to his asymptomatic status while in Tianjin, and then infected the case through frequent close contact at work.
Furthermore, COVID-19 virus RT-PCR test results on throat, nasopharyngeal, and anal swab samples were all positive with cycle threshold (Ct) values of 16.0/18.0 (ORF1ab/N), 23.0/24.4, and 35.0/35.0, respectively. As the Ct values of the anal swab sample was too high to acquire whole genome sequences, we chose throat (FH-124-Y) and nasopharyngeal (FH-124-B) swab samples for genome sequencing. The whole genome of FH-124-Y was 29,868 nt, and FH-124-B was 29,803 nt, with a shortage in 5’-NTR and 3’-polyA tail. The genome sequences of FH-124-Y and FH-124-B were found to be 100% identical and were both named Tianjin/137/2020.
Phylogenetic analysis has revealed that all known COVID-19 virus genomes fall into two major clades, or lineages, named S and L (Figure 1). We found that Tianjin/137/2020 belonged to the Europe Branch of the L-Lineage, and that it was most closely related to the Beijing strain ESP_ISL_469255 (4), to which it was 100% identical at the nucleotide level.
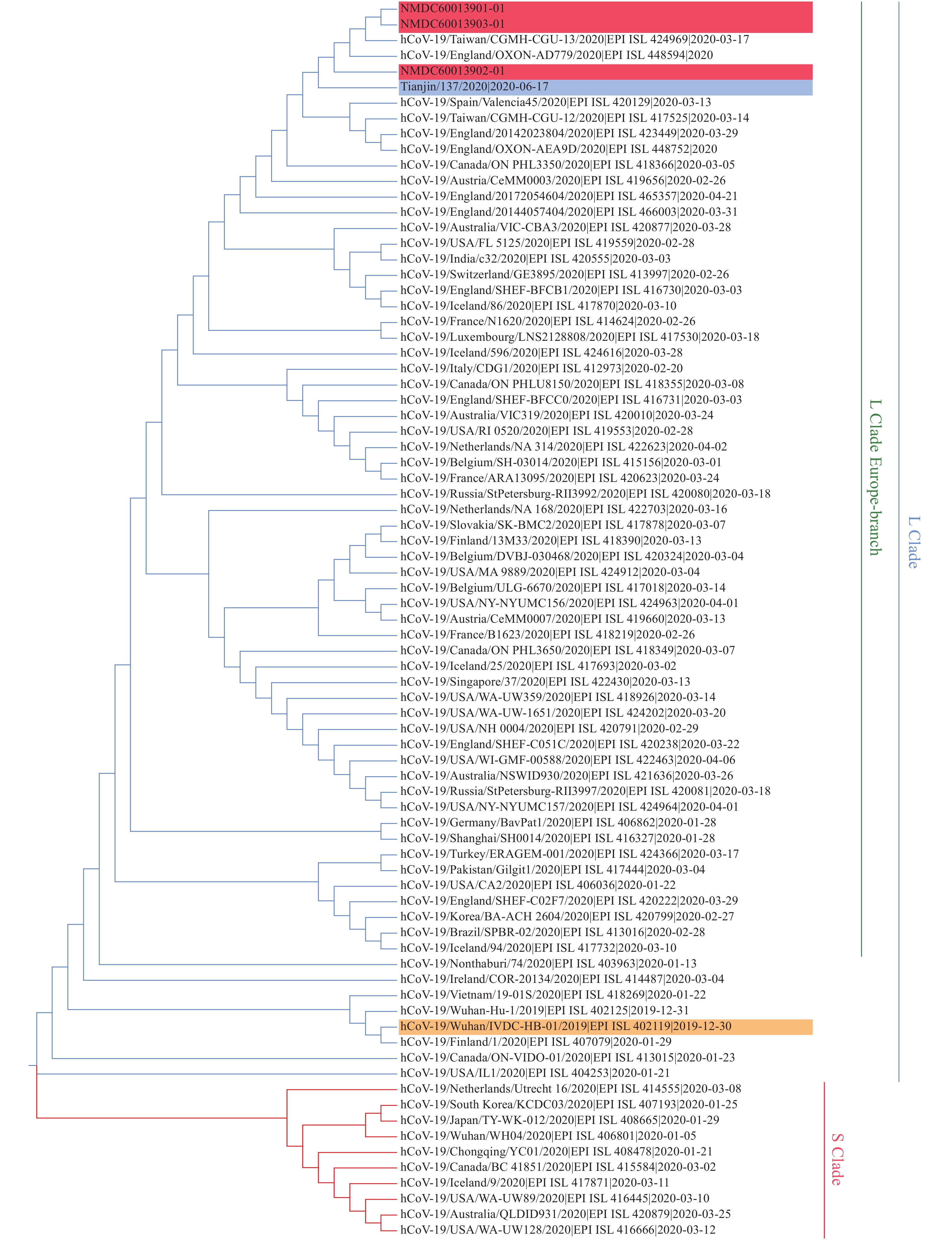 Figure 1.
Figure 1.Phylogenetic tree based on the whole genome sequences of COVID-19 virus. The COVID-19 virus genome of the cases in Beijing Municipality and Tianjin City were highlighted in shades of red and blue, respectively. The reference sequence Wuhan-Hu-1 (NC_045512) were highlighted in orange. S- or L-clade of COVID-19 virus were marked and colored on the right.
-
The 137th patient had no contact with individuals who arrived in Tianjin from abroad or other PLADs before illness onset. Foreigners who stayed in the hotel where the patient had worked were isolated for 14 days; their tests for COVID-19 virus nucleic acids and serum antibodies were all negative. Closed-loop isolation management was implemented for all foreign personnel, and the possibility of the patient acquiring infection from these individuals was excluded.
No new local cases were reported in Tianjin for over 110 days. Family members, colleagues, neighbors, and hotel guests were investigated. None of them had any suspicious symptoms, and the results of nucleic acid and serum antibody tests were negative. Therefore, the possibility that other unknown local cases were the source was excluded.
Local CDC professionals collected hundreds of specimens from surfaces in the patient’s residence and workplace and from meat, seafood, and other food products in the hotel. They also collected environmental and product-related samples from the hotel’s supplier. All samples were subjected to nucleic acid testing and all results were negative, thereby excluding the possibility that surface or food contamination was the source.
The serological test result of 1 of 55 hotel dining room employees was positive for COVID-19-virus-specific IgM antibody. This individual worked as a chef in the western dining room of the hotel but had reported no COVID-19-like symptoms. The chef had been to Beijing many times in the past month to see his girlfriend. In particular, he visited restaurants and bars, a theater, and other public places in Beijing on June 8. He had close communication with the patient on June 10, during which, the chef did not wear a mask. During the typical work day, the two men met in the employees’ dining room during the daily lunch and often smoked together in the smoking room on the ground floor of the hotel several times.
COVID-19 virus genomes fall into two major types (L and S), defined by two different SNPs at nucleotide 8,782 and 28,144. Strains that exhibit a “CT” haplotype at these two sites are designated as L type and strains that exhibit a “TC” haplotype are designated as S type. Although the L type is more prevalent (≥70%) than the S type (≥30%), the S type is evolutionarily older (5).
Our phylogenetic tree clearly showed the separation of the 2 lineages. Tianjin/137/2020 and ESP_ISL_469255 (from the Beijing outbreak) clustered together in the L-Lineage of the Europe-Branch. Beijing strains ESP_ISL_469254, ESP_ISL_469256, Taiwan strain CGMH-CGU-13, and England strain OXON-AD779, were also found tightly clustered, indicating that Tianjin/137/2020 was closely related to the Beijing and Europe strains rather than local strains identified during the first wave of the epidemic in China, which initiated the epidemic in Wuhan.
Evidence from the epidemiological investigations, nucleic acid and antibody tests, and whole genome sequencing indicated that the chef was most likely infected with COVID-19 virus in public places in Beijing. The chef was asymptomatic and went back to work, transmitting the virus to the 137th patient. Therefore, we believe that this patient in Tianjin is linked to the Xinfadi Agricultural Product Wholesale Market in Beijing by traditional and molecular epidemiology.
HTML
| Citation: |



 Download:
Download:




